Genetic Diversity of Malaysia
Background
In the early 1990s, we collaborated with Dr. Yik-Yuen Gan, to study genetic diversity in Malaysia. She was working in the Department of Biotechnology at the Universiti Pertanian Malaysia at the time and had come to Emory University to learn mitochondrial genetics in the laboratory of Dr. Douglas Wallace. Through her engagement with local communities, we analyzed DNA samples from Malaysian Han Chinese individuals originating in the Fujian/Guangdong region of China, ethnic Malays, Orang Asli individuals, and indigenous Borneans from Sabah State (Borneo).
In the following sections, we will discuss the analysis of mtDNA variation in Malaysian populations, which was conducted as part of a larger analysis of genetic diversity in Southeast Asian populations. While a fuller description of this work appears in our Schurr and Wallace (2002) paper – which is discussed in a separate webpage on Southeast Asian mtDNA diversity – we will focus on Malaysian population diversity.
To further contextualize this research, we will draw upon information from several other papers that have since expanded our understanding of Malaysian population diversity.
A Map of Malaysia. Source: https://www.thoughtco.com/faq-about-singapore-195082
History of Orang Asli and Malays Tribes in Malaysia
History of Orang Asli and Malays Tribes in Malaysia Text
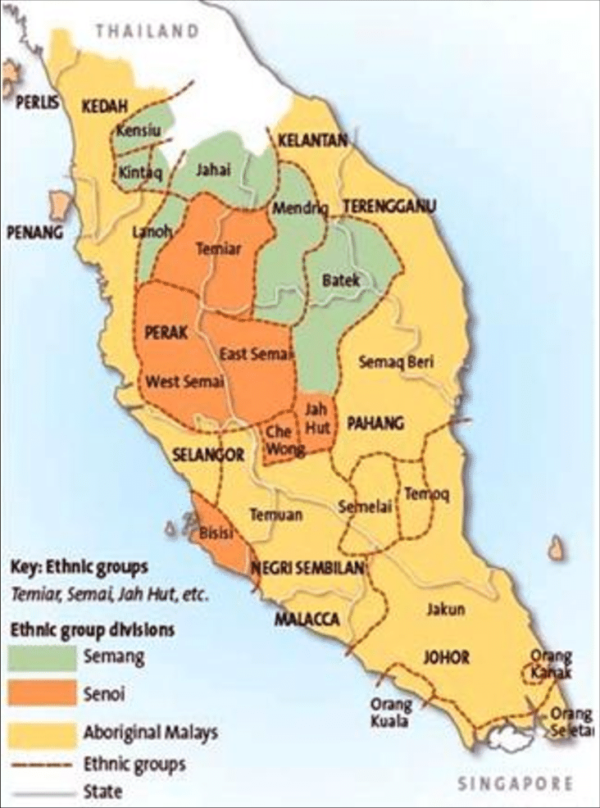
Map of Orang Asli and Malays Tribes in Malaysia. Source: XXXX
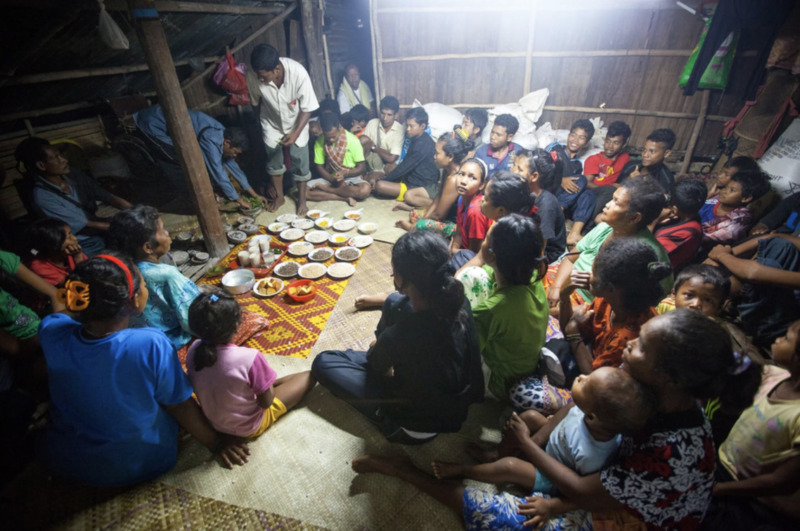

A family gathering inside an Orang Asli home. Photo credit: © De Visu / Shutterstock
History of Indigenous Populations of Borneo
History of Indigenous Populations of Borneo texts
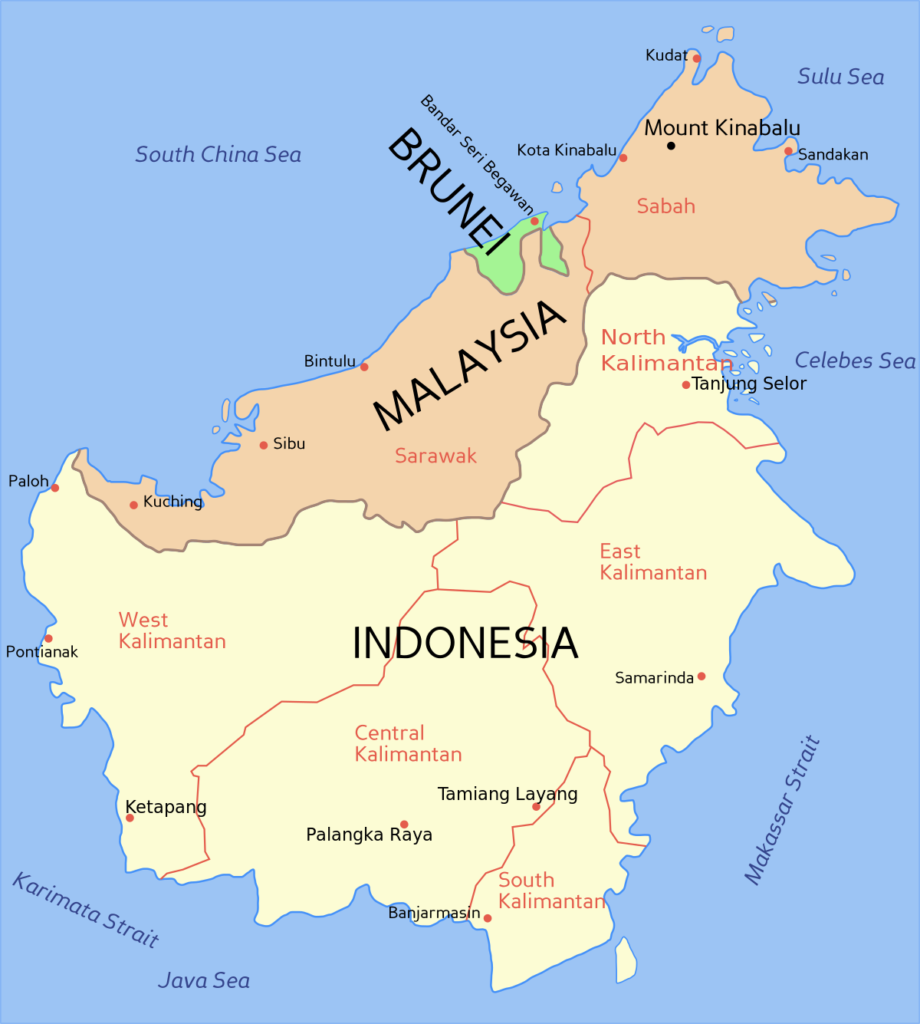
Map of Borneo. The political subdivisions of the island in Malaysia and Indonesia are shown. Source: XXXXX
Genetic Diversity in Malaysian Populations
In the following discussion, we will discuss the analysis of mtDNA variation in Malaysian populations conducted in the early 1990s as part of a larger analysis of genetic diversity in Southeast Asian populations. While a fuller description of this work appears in our Schurr and Wallace (2002) paper – which is discussed in a separate webpage on Southeast Asian mtDNA diversity – this discussion focuses on Malaysian and the different ethnic groups living there.
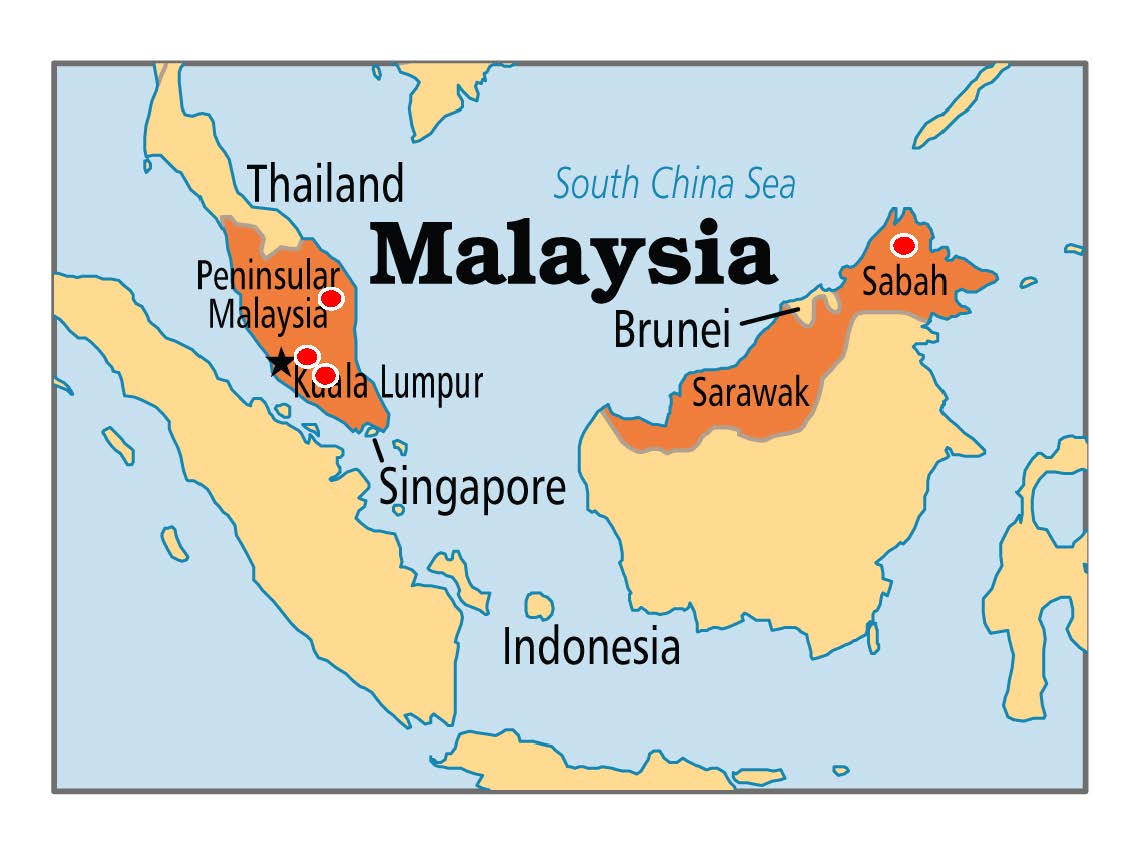
Map of Southeast Asia Showing the Sample Localities for Malaysian and Southeast Asian Populations Studied for mtDNA Diversity. Map Source: https://babogenglish.wordpress.com/2016/07/27/malaysia-general-information/
To analyze sequence variation in each mtDNA, we PCR amplified the entire mitogenome sequence in nine overlapping fragments and subjected the resulting amplicons to restriction fragment length polymorphism analysis using 18 endonucleases (AluI, AvaII, BamHI, DdeI, HaeII, HaeIII, HhaI, HincII, Hinfl, HpaI, HpaII, MboI, Pstl, PvuII, RsaI, TaqI, XbaI, XhoII). In addition, each sample was screened for the COII/tRNA-Lys length mutations (see Ballinger et al. 1992 for details). The fragments detected through this method were mapped against the published mtDNA sequence (Anderson et al. 1981) and the positions of variants were numerically assigned based on the restriction site maps predicted for each endonuclease. The variants identified in each sample were then combined to define its haplotype. Sets of related haplotypes that shared a common set of variants were called haplotype groups (now haplogroups).
The table below present the haplotype groups observed in Southeast Asian populations along with a detailed discussion of their derivation from mtDNA RFLP data.
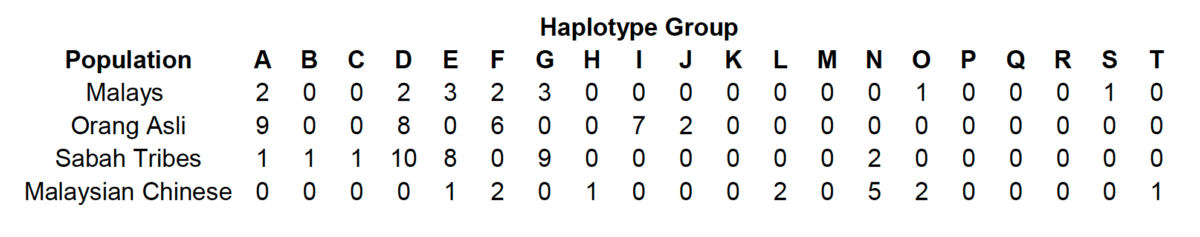
Haplotype groups were classified according to polymorphic sites present in Malaysian mtDNAs.
In what appears below, the restriction site gains and losses for each haplotype group relative to the published sequence (Anderson et al. 1981) are shown in bold face and non-bold face, respectively; slashes between enzyme letters or sites indicate non-independent events. Restriction sites enclosed in brackets indicate sites that frequently accompany the definitive sites for a haplotype group. The letter designation of restriction enzymes are as follows: a, AluI; b, Avall; c, DdeI; e, HaeIII; f, HhaI; g, Hinfl; h, Hpal; i, HpaII; j, MboI; k, RsaI; I, TaqI; m, BamHI; n, HaeII; 0, HinclI. The COII/tRNA-Lys 9-bp intergenic deletion occurs within haplotype groups A-D and F; the 4-bp insertion occurs within haplotype groups A and M.
Haplotype groups are indicated by capital letters and consist of sets of polymorphic restriction sites; sites shown in brackets occur frequently but not exclusively in a particular haplotype group:
[A] 12406h/12406o, 16517e [9052n/9053f];
[B] 7853o, 10394c, 10397a;
[C] 3534c/3537a, 10394c, 15234g/15235j, 16517e;
[D] 16517e;
[E] 10394c, 10397a, 16517e;
[F] 10394c, 10397a;
[G] 7598f, 10394c, 10397a, 16517e, 16389g/16390b;
[H] 663e [16517e];
[1] 1063e, 9326n/9329f, 10394c, 10397a; 10143a [951j;
[J] 1715c, 4711i, 10394c, 11403g. 131180j;
[K] 4830n/4831f, 10394c, 10397a;
[L] 5176a, 10394c, 10397a;
[M] 10394c [16517e];
[N] 16517e;
[O] 13366b/l3367j/l3368m [16517e];
[P] 12026h/12026o;
[Q] 1002q/1004o;
[R] 10394c, 10397a, 13259o/13261a, 16517e;
[S] 207h/207o, 15606a;
[T] 16389g/16390b, 16517e.
Based on updated information about the haplotype groupings, we can change the designations some of these maternal lineages to reflect their currently understood mutational properties and corresponding haplogroups. In other cases, the original RFLP data were not granular enough to allow for specific translations of the 1992 data into modern lineage designations. As a result, those haplotype groups are assigned to their putative macrohaplogroup status (M, N, R) (see also Schurr and Wallace 2002).
[A] 12406h/124060, 16517e, [9052n/9053f] => F
[B] 7853o, 10394c, 10397a => M
[C] 3534c/3537a, 10394c, 15234g/15235j, 16517e => B5
[D} 9-bp deletion, 16517e => B
[E] 10394c, 10397a, 16517e => M
[F] 10394c, 10397a => M
[G] 7598f, 10394c, 10397a, 16389g/16390b, 16517e => E
[H} 663e [16517e] => A
[1] 9326n/9329f, 10394c, 10397a, 10143a [951j, 1063e] => M
[J] 1715c, 4711i, 10394a, 11403g. 131180j => N
[K] 4830n/4831f, 10394c, 10397a => G
[L] 5176a, 10394c, 10397a => D
[M] 10394c [16517e] => N
[N] 16517e => R
[O] 13366b/13367j/13368m [16517e] => R
[P] 12026h/12026o => R
[Q] 1002q/1004o => R
[R] 13259o/13261a, 10394c, 10397a, 16517e => C
[S] 207h/207o, 15606a => P7
[T] 16389g/16390b, 16517e => R
The table below present the mtDNA data observed in Malaysian populations based on updated haplogroup nomenclature.
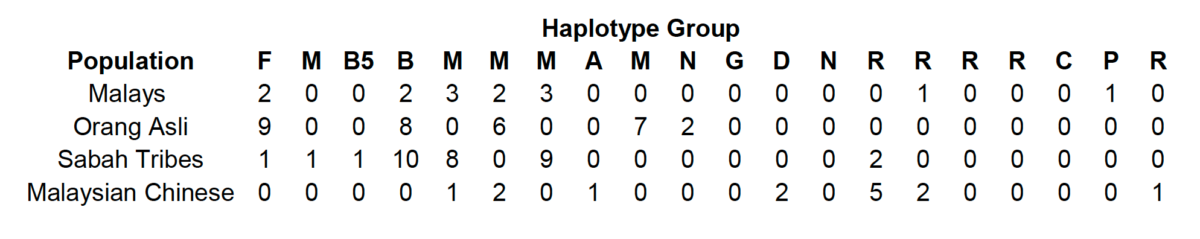
Although we don’t know the exact classificatory statuses of some of the M-, N- and R-derived haplogroup in Malaysian populations, it is clear that they are somewhat genetically distinctive relative to Malaysian Han Chinese, who are more recent immigrants to the region.
Subsequent studies of Malaysian populations by other scholars employed control region sequencing of the mtDNA along with targeted SNP genotyping to delineate maternal lineages present in them (Hill et al. 2006, 2007). This approach provided a more fine-grained view of mtDNA diversity in Malaysian populations than possible to generate using only RFLP data. In addition, some of the RRLPs in our study identified coding region variants not detectable through control region sequencing, making an exact comparison of the haplogroup datasets complicated.
Below, a schematic tree of East Eurasian mtDNA haplogroups, displaying the various branches of macrohaplogroup M (A) and macrohaplogroup N (B) and indicating diagnostic control-region positions and coding-region positions appearing in different haplogroups. R1270 has been renamed as R22, since R12 was used in a previous study. F1a3 is here defined solely by 16311 because the two other control-region sites (Kong et al. 2006) did not appear in these samples (from Hill et al. 2007).
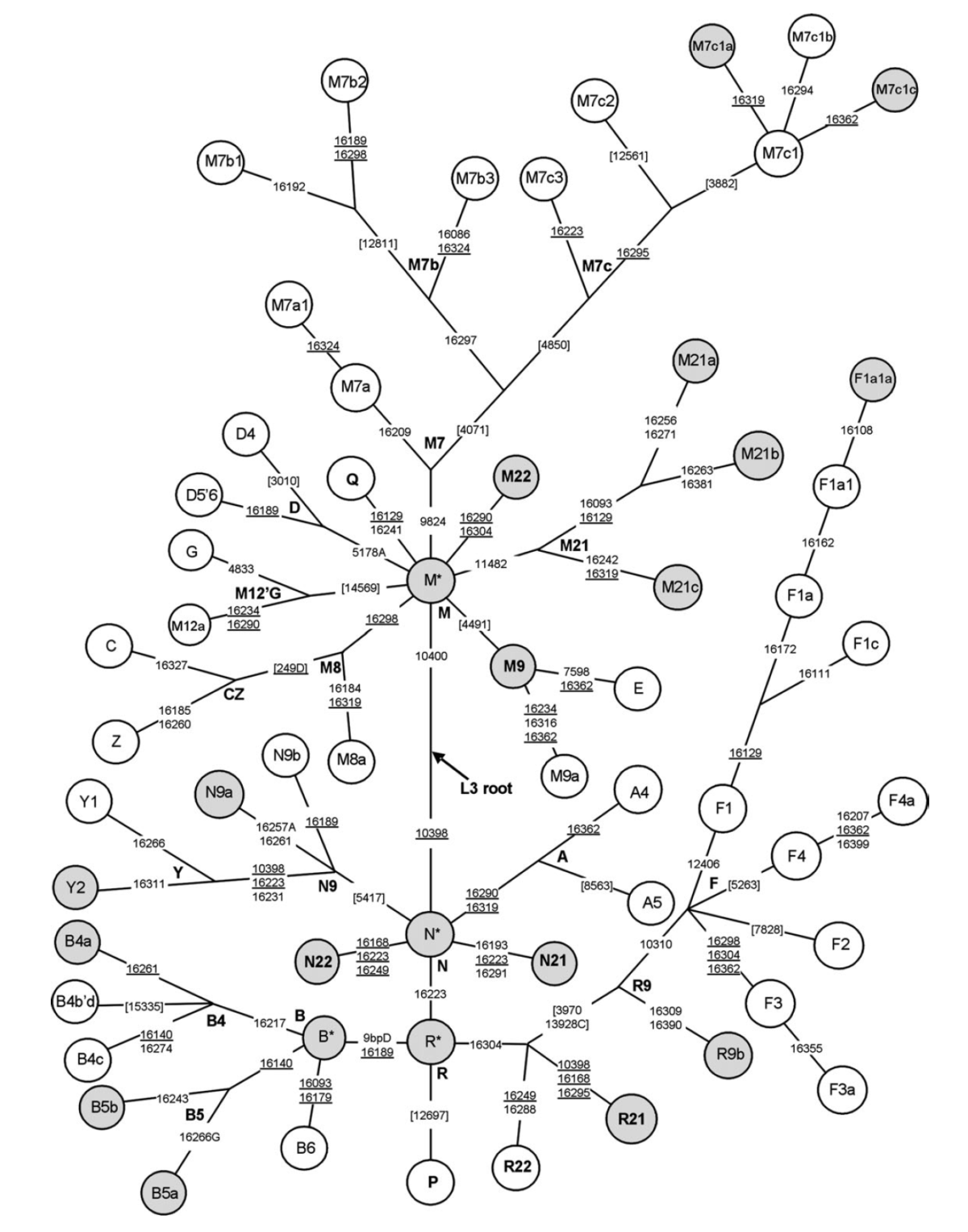
Schematic tree of East Eurasian mtDNA haplogroups, displaying the various branches of macrohaplogroup M (A) and macrohaplogroup N (B) and indicating diagnostic control-region positions and coding-region positions tested. Here, R1270 has been renamed as R22, since R12 was used in a previous study. F1a3 is here defined solely by 16311 because the two other control-region sites (Kong et al. 2006) do not appear in these samples (from Hill et al. 2007).
Publication
Ballinger SW, Schurr TG, Torroni A, Gan YY, Hodge JA, Hassan K, Chen K-H, Wallace DC. 1992. Southeast Asian mitochondrial DNA analysis reveals genetic continuity of ancient Mongoloid migrations. Genetics 130(1):139-152. Link
Hill C, et al. 2006. Phylogeography and ethnogenesis of aboriginal Southeast Asians. Mol. Biol. Evol. 23(12):2480–2491;
Hill et al. 2007. A mitochondrial stratigraphy for island Southeast Asia. Am J Hum Genet 80:29-43.
More Information
sdafsadfgj queruqweut dadf
Laboratory of Molecular Anthropology
University of Pennsylvania
421 University Museum
3260 South Street
Philadelphia, PA 19104-6398
Tel: 215-573-2656

FOLOW US:
